Molecular docking is a computer simulation technique to predict drug interactions and biological molecules. This article provides an overview of the basic principles, methodologies, and specific applications of molecular docking, focusing on the HDock method.
Understanding this enables one to effectively screen and design drugs for new drug development and protein research. It also helps in elucidating protein interactions and optimizing drug molecules. Furthermore, integrating knowledge of computational science with life sciences allows for deeper understanding and application.
Give it a try!
macOS Ventura(13.2.1)
To proceed with this article, please download PyMOL.
What is Molecular Docking?
Molecular docking is a computer simulation predicting how drugs or biological molecules (e.g., proteins) will interact. It calculates how a specific protein and drug will bind, which parts touch each other, and predicts the drug’s effects based on these interactions. This is extremely beneficial for new drug development. Numerous web-based molecular docking methods are available. This article will introduce HDock, which handles protein-protein docking.
Protein Preparation
First, let’s prepare the protein for docking before analyzing each method. For this example, we will use the leptin protein and its receptor with the Protein Data Bank (PDB) number 8DHA.
Leptin is a protein hormone derived from fat tissue, playing a crucial role in regulating weight and energy balance. It’s released from fat cells into the bloodstream and binds to specific receptors in the hypothalamus, suppressing appetite and regulating metabolism.
Leptin is a protein hormone derived from fat tissue, playing a crucial role in regulating weight and energy balance. It’s released from fat cells into the bloodstream and binds to specific receptors in the hypothalamus, suppressing appetite and regulating metabolism.
The appeal of leptin drugs is their potential to suppress appetite and induce weight loss. This could aid weight management and reduce disease risks for obese or metabolic syndrome patients. Additionally, as leptin is involved in neural circuits controlling appetite, there’s potential for developing new appetite-controlling therapies.
Recent success includes lowering blood sugar using leptin without insulin, offering promise for diabetes treatment.
The leptin and its receptor with the PDB number 8DHA represent only the binding region where they are connected, and the entire structure is registered as 8DH8. To reduce computational complexity in this case, we will utilize the leptin and its receptor with the PDB number 8DHA.
In Pymol, choose File → Get PDB…, enter 8DHA, and display the leptin and its receptor.
Next, use Display → Sequence to show the full sequence.
Select the sequence starting from /C/C/1, which corresponds to leptin.
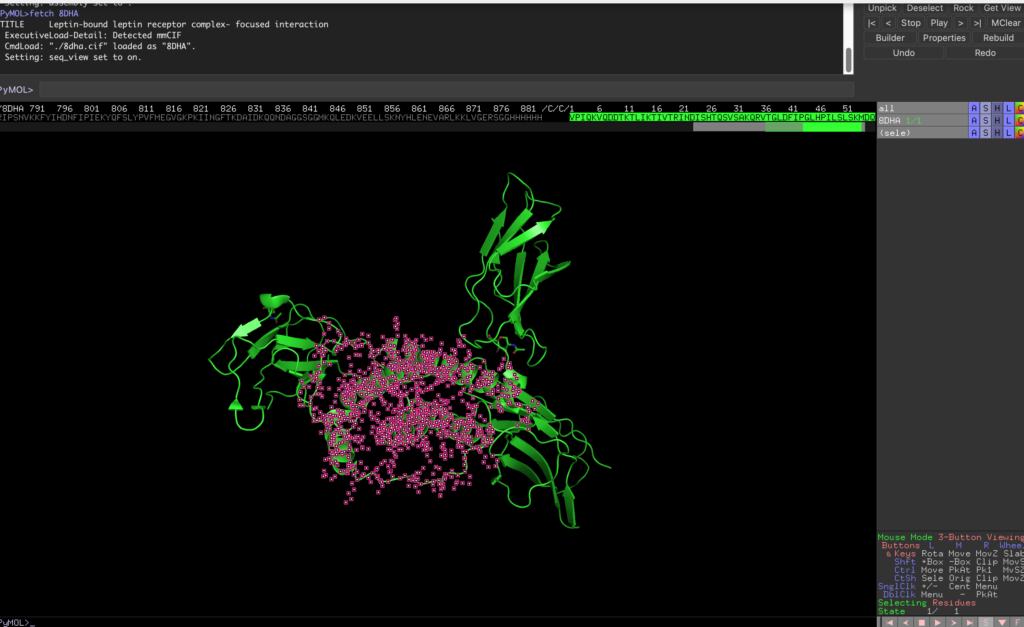
Then, execute the following code to save the selection as leptin.pdb:
save leptin.pdb, seleSave the receptor part displayed as focus_leptinR.pdb:
save focus_leptinR.pdb, seleAvoid selecting 2-acetamido-2-deoxy-beta-D-glucopyranose.
These files should be saved in Macintosh HD/Users/Your Computer Name.
Protein preparation is now complete.
What is HDock?
HDock works by exploring possible bindings between a ligand and a receptor (docking) and then evaluating the quality of these bindings (scoring). HDock can predict the most natural binding between the ligand and the receptor. HDock is a web server designed not just for protein-protein but also for protein-DNA/RNA docking. We will be trying out protein-protein docking in this instance.
Let’s try HDock right away!
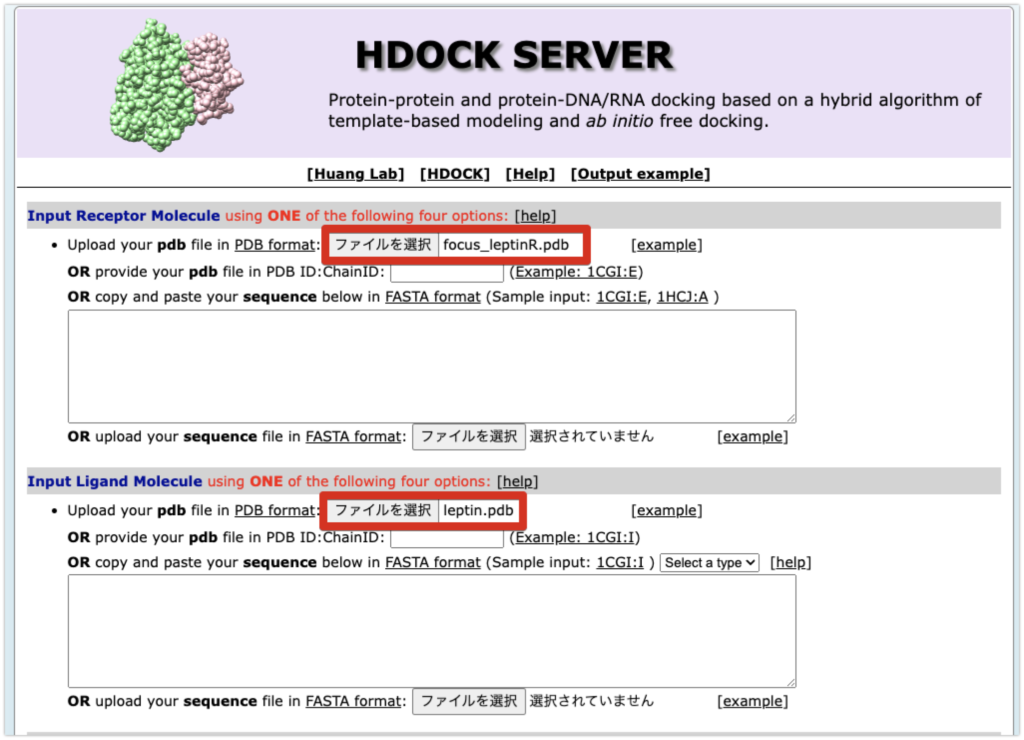
Please go to the HDock site and enter the following:
- In “Input Receptor Molecule”: focus_leptinR.pdb
- In “Input Ligand Molecule”: leptin.pdb
- Please input your email address and job name.
- Click “Submit” and wait for a while. The results will be available in about two hours.
Mechanism of HDock
Let’s delve deeper into the mechanism of HDock. HDock mainly operates in two phases: the ‘docking’ phase, which generates potential bindings between the ligand and receptor, and the ‘scoring’ phase, which evaluates the quality of these bindings.
- Docking Phase: HDock explores all possible bindings between the ligand and receptor, changing the structure of the ligand to see how it fits with the receptor. Numerous potential bindings are generated in this phase.
- Scoring Phase: This phase evaluates the quality of the generated bindings. HDock assesses how ‘natural’ each binding is or how likely it would occur in a real biological environment. This is done by calculating binding energies. The most natural binding (i.e., the binding with the lowest energy) is considered the best.
Result
Here are the results (they seem to be stored on the server for about two weeks, so download them to your computer).
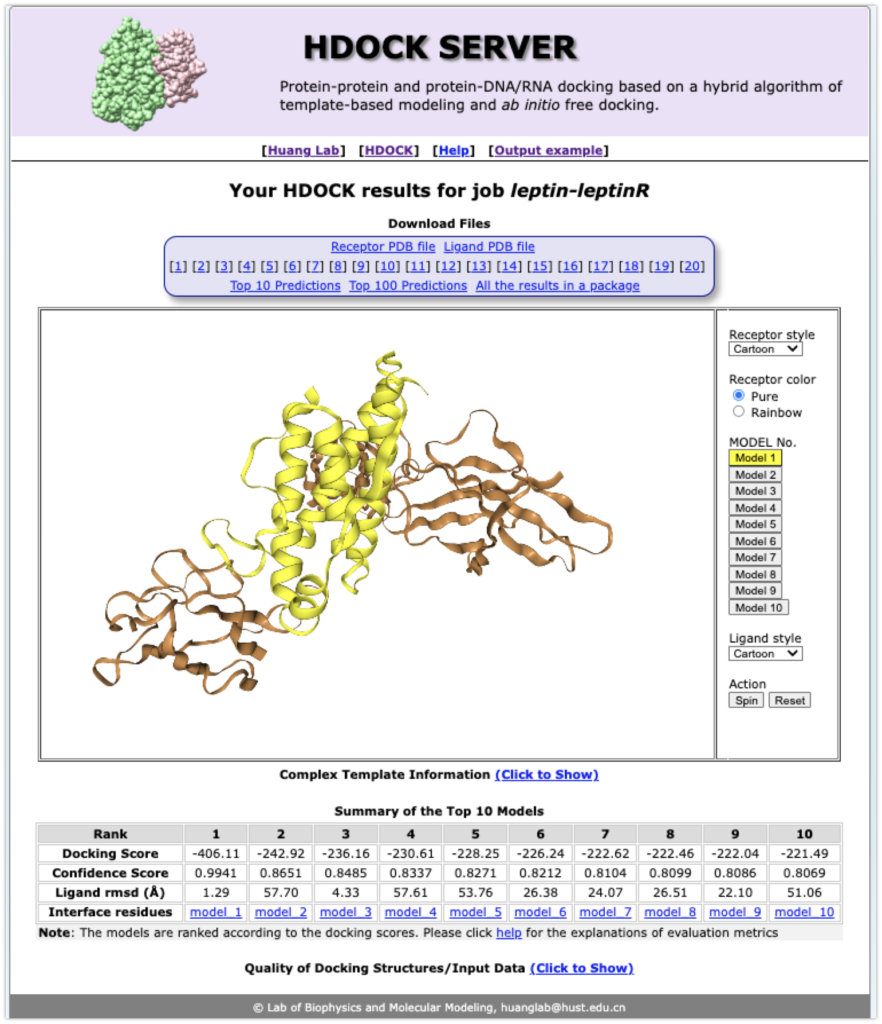
Display the model with the best docking score in PyMOL.
The colors have been slightly altered: leptin.pdb is flesh-colored, and focus_leptinR.pdb is green.
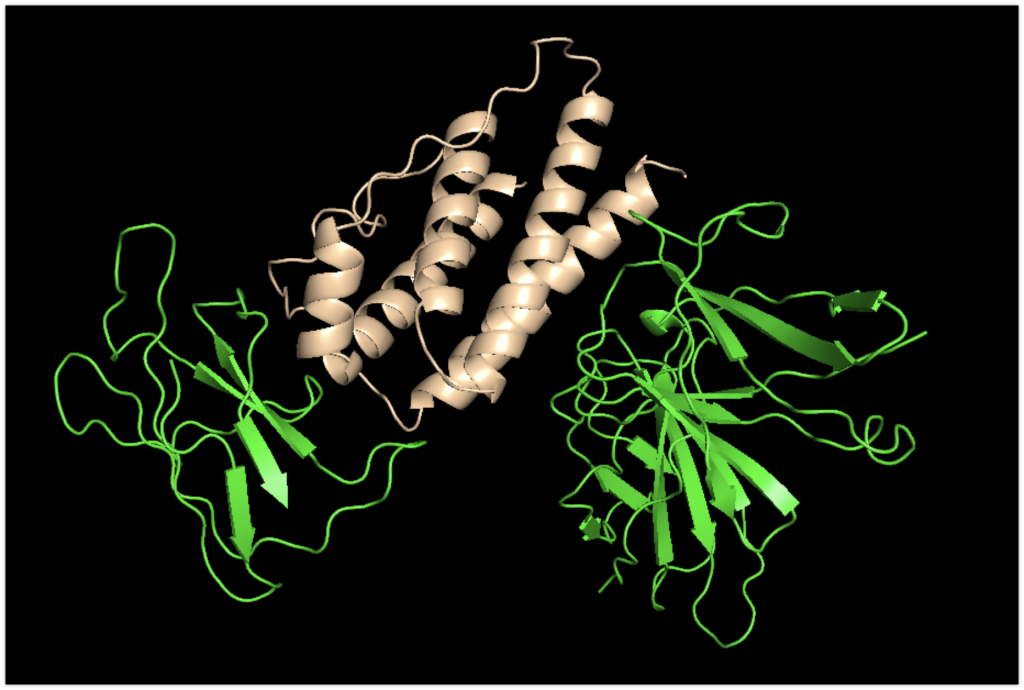
Now, let’s compare it with the original sequence.
Go to the PyMOL tab, File → Get PDB… and type in “8DHA”.
The magenta structure represents the original complex structure recorded in the PDB.
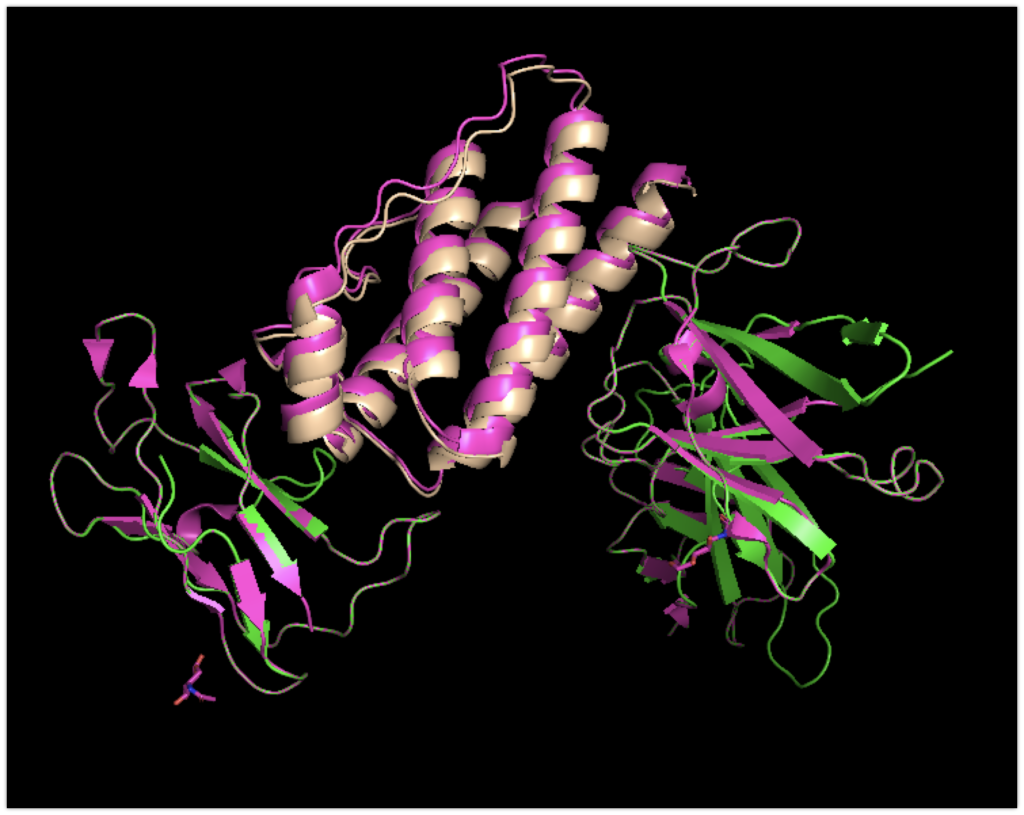
While there are slight deviations, it’s evident that it binds with good accuracy.
In Conclusion
How was it? Today, we explored HDock among protein-protein docking methods. Since you can easily try docking on this server, I encourage everyone to try it. Previous articles covered ClusPro, PatchDock, and LZerD pairwise docking. HDock can dock with high accuracy with the proteins used this time.
HDock can also dock DNA with proteins.
Furthermore, you can dock using only sequences, making it very convenient.
Do try these methods!

